What you need to know
-
SARS-CoV-2 is genetically similar to SARS-CoV-1, but characteristics of SARS-CoV-2—eg, structural differences in its surface proteins and viral load kinetics—may help explain its enhanced rate of transmission
-
In the respiratory tract, peak SARS-CoV-2 load is observed at the time of symptom onset or in the first week of illness, with subsequent decline thereafter indicating the highest infectiousness potential just before or within the first five days of symptom onset
-
Reverse transcription polymerase chain reaction (RT-PCR) tests can detect viral SARS-CoV-2 RNA in the upper respiratory tract for a mean of 17 days; however, detection of viral RNA does not necessarily equate to infectiousness, and viral culture from PCR positive upper respiratory tract samples has been rarely positive beyond nine days of illness
-
Symptomatic and pre-symptomatic transmission (1-2 days before symptom onset), is likely to play a greater role in the spread of SARS-CoV-2 than asymptomatic transmission
-
A wide range of virus-neutralising antibodies have been reported, and emerging evidence suggests that these may correlate with severity of illness but wane over time
Since the emergence of SARS-CoV-2 in December 2019, there has been an unparalleled global effort to characterise the virus and the clinical course of disease. Coronavirus disease 2019 (covid-19), caused by SARS-CoV-2, follows a biphasic pattern of illness that likely results from the combination of an early viral response phase and an inflammatory second phase. Most clinical presentations are mild, and the typical pattern of covid-19 more resembles an influenza-like illness—which includes fever, cough, malaise, myalgia, headache, and taste and smell disturbance—rather than severe pneumonia (although emerging evidence about long term consequences is yet to be understood in detail).1 In this review, we provide a broad update on the emerging understanding of SARS-CoV-2 pathophysiology, including virology, transmission dynamics, and the immune response to the virus. Any of the mechanisms and assumptions discussed in the article and in our understanding of covid-19 may be revised as further evidence emerges.
What we know about the virus
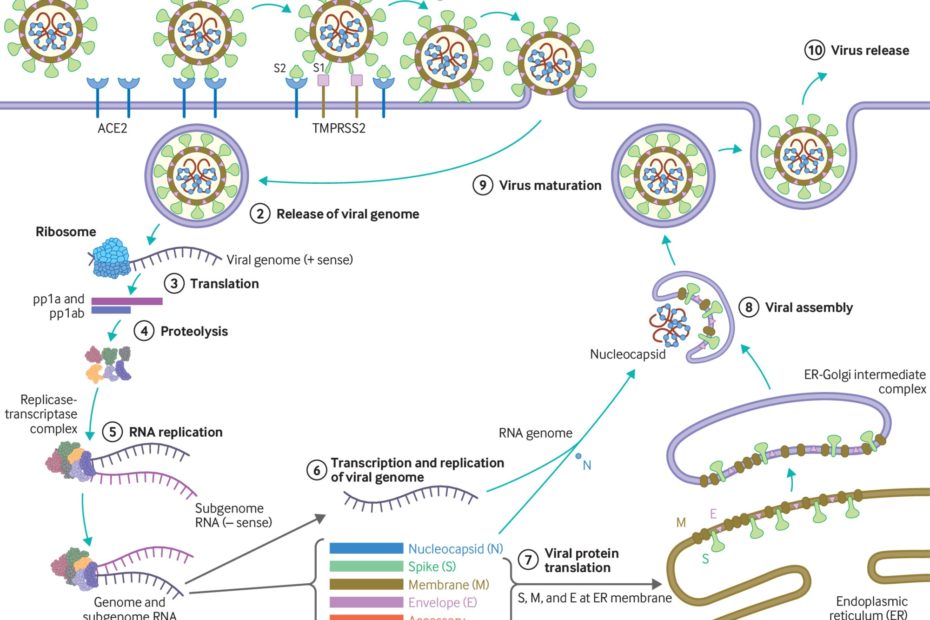
SARS-CoV-2 is an enveloped β-coronavirus, with a genetic sequence very similar to SARS-CoV-1 (80%) and bat coronavirus RaTG13 (96.2%).2 The viral envelope is coated by spike (S) glycoprotein, envelope (E), and membrane (M) proteins (fig 1). Host cell binding and entry are mediated by the S protein. The first step in infection is virus binding to a host cell through its target receptor. The S1 sub-unit of the S protein contains the receptor binding domain that binds to the peptidase domain of angiotensin-converting enzyme 2 (ACE 2). In SARS-CoV-2 the S2 sub-unit is highly preserved and is considered a potential antiviral target. The virus structure and replication cycle are described in figure 1.
Fig 1
(1) The virus binds to ACE 2 as the host target cell receptor in synergy with the host’s transmembrane serine protease 2 (cell surface protein), which is principally expressed in the airway epithelial cells and vascular endothelial cells. This leads to membrane fusion and releases the viral genome into the host cytoplasm (2). Stages (3-7) show the remaining steps of viral replication, leading to viral assembly, maturation, and virus release
Coronaviruses have the capacity for proofreading during replication, and therefore mutation rates are lower than in other RNA viruses. As SARS-CoV-2 has spread globally it has, like other viruses, accumulated some mutations in the viral genome, which contains geographic signatures. Researchers have examined these mutations to study virus characterisation and understand epidemiology and transmission patterns. In general, the mutations have not been attributed to phenotypic changes affecting viral transmissibility or pathogenicity. The G614 variant in the S protein has been postulated to increase infectivity and transmissibility of the virus.3 Higher viral loads were reported in clinical samples with virus containing G614 than previously circulating variant D614, although no association was made with severity of illness as measured by hospitalisation outcomes.3 These findings have yet to be confirmed with regards to natural infection.
Why is SARS-CoV-2 more infectious than SARS-CoV-1?
SARS-CoV-2 has a higher reproductive number (R0) than SARS-CoV-1, indicating much more efficient spread.1 Several characteristics of SARS-CoV-2 may help explain this enhanced transmission. While both SARS-CoV-1 and SARS-CoV-2 preferentially interact with the angiotensin-converting enzyme 2 (ACE 2) receptor, SARS-CoV-2 has structural differences in its surface proteins that enable stronger binding to the ACE 2 receptor4 and greater efficiency at invading host cells.1 SARS-CoV-2 also has greater affinity (or bonding) for the upper respiratory tract and conjunctiva,5 thus can infect the upper respiratory tract and conduct airways more easily.6
Viral load dynamics and duration of infectiousness
Viral load kinetics could also explain some of the differences between SARS-CoV-2 and SARS-CoV-1. In the respiratory tract, peak SARS-CoV-2 load is observed at the time of symptom onset or in the first week of illness with subsequent decline thereafter, which indicates the highest infectiousness potential just before or within the first five days of symptom onset (fig 2).7 In contrast, in SARS-CoV-1 the highest viral loads were detected in the upper respiratory tract in the second week of illness, which explains its minimal contagiousness in the first week after symptom onset, enabling early case detection in the community.7
Fig 2
After the initial exposure, patients typically develop symptoms within 5-6 days (incubation period). SARS-CoV-2 generates a diverse range of clinical manifestations, ranging from mild infection to severe disease accompanied by high mortality. In patients with mild infection, initial host immune response is capable of controlling the infection. In severe disease, excessive immune response leads to organ damage, intensive care admission, or death. The viral load peaks in the first week of infection, declines thereafter gradually, while the antibody response gradually increases and is often detectable by day 14 (figure adapted with permission from doi:10.1016/j.cell.2020.04.013; doi:10.1016/S2213-2600(20)30230-7)
Quantitative reverse transcription polymerase chain reaction (qRT-PCR) technology can detect viral SARS-CoV-2 RNA in the upper respiratory tract for a mean of 17 days (maximum 83 days) after symptom onset.7 However, detection of viral RNA by qRT-PCR does not necessarily equate to infectiousness, and viral culture from PCR positive upper respiratory tract samples has been rarely positive beyond nine days of illness.5 This corresponds to what is known about transmission based on contact tracing studies, which is that transmission capacity is maximal in the first week of illness, and that transmission after this period has not been documented.8 Severely ill or immune-compromised patients may have relatively prolonged virus shedding, and some patients may have intermittent RNA shedding; however, low level results close to the detection limit may not constitute infectious viral particles. While asymptomatic individuals (those with no symptoms throughout the infection) can transmit the infection, their relative degree of infectiousness seems to be limited.91011 People with mild symptoms (paucisymptomatic) and those whose symptom have not yet appeared still carry large amounts of virus in the upper respiratory tract, which might contribute to the easy and rapid spread of SARS-CoV-2.7 Symptomatic and pre-symptomatic transmission (one to two days before symptom onset) is likely to play a greater role in the spread of SARS-CoV-2.1012 A combination of preventive measures, such as physical distancing and testing, tracing, and self-isolation, continue to be needed. (…)
- Muge Cevik, clinical lecturer1 2,
- Krutika Kuppalli, assistant professor3,
- Jason Kindrachuk, assistant professor of virology4,
- Malik Peiris, professor of virology5
1Division of Infection and Global Health Research, School of Medicine, University of St Andrews, St Andrews, UK
2Specialist Virology Laboratory, Royal Infirmary of Edinburgh, Edinburgh, UK and Regional Infectious Diseases Unit, Western General Hospital, Edinburgh, UK
3Division of Infectious Diseases, Medical University of South Carolina, Charleston, SC, USA
4Laboratory of Emerging and Re-Emerging Viruses, Department of Medical Microbiology, University of Manitoba, Winnipeg, MB, Canada
5School of Public Health, LKS Faculty of Medicine, The University of Hong Kong, Hong Kong Special Administrative Region, China
- Correspondence to M Cevik mc349{at}st-andrews.ac.uk